https://doi.org/10.3390/molecules27238194
”
3.2. Preparation of Standard Solutions
Stock standard solutions (25 mg/mL) were prepared by weighing the appropriate amount of each standard into a 20 mL volumetric flask and dissolving in water. Mixed standard solution I (2.5 mg/mL) was prepared by water dilution of 5 mL of each stock standard solutions in 50 mL volumetric flask. Mixed stock solution II was prepared from mixed stock solution I by dilution (1/10) with water to yield a concentration of 0.25 mg/mL. Working solutions were prepared by dilution of required aliquots of the mixed standard solution I or II. Internal standard solution (5 mg/mL) was prepared by dissolving 250 mg 1,7-diaminoheptane in 50 mL of water. All solutions were stored at 2–9 °C.
3.3. Apparatus and Chromatographic Conditions
The instrumental analysis was performed using HPLC-DAD Varian Model 330 Pro Star (Varian, The Netherlands) equipped with ternary pump, autosampler and column thermostat, controlled by the Galaxie Workstation software. Separation was conducted on Unisol C18 150 × 4.6 mm, 3 μm with precolumn 10 × 3 mm, 3 μm (Agela Technologies, Wilmington, DE, USA). The column oven temperature was maintained at 30 °C and the analytes were separated using acetonitrile (mobile phase A) and water (mobile phase B). The flow rate of 1 mL/min was used. Gradient program was made as follows: 60% (mobile phase A) increased to 75% in 6 min and hold 2 min, increased to 95% in 5 min and hold 7 min, decreased to 60% in 1 min and hold with 9 min to column recondition. Total analysis run time was 30 min, and the injection volume was 10 μL.
3.4. Sample Preparation
Five grams of homogenized cheese sample was weighted in a 50 mL centrifuge tube and 100 μL of the internal standard (5 mg/mL, 1,7-diaminoheptane) was added. Then, ceramic homogenizer and 10 mL of 0.2 M hydrochloric acid were added and sample was shaken for 1 min and in an Analog Orbital Shaker (Sunlab Instruments, Germany) for 5 min. The obtained cheese slurry was centrifuged at 3000 rpm for 20 min at 4 °C and the upper fat layer was removed. In case of unclear supernatants, they were filtered through a PTFE filter. After that, 100 μL was transferred into a screw cap glass vial and 200 μL of saturated sodium carbonate solution and 400 μL of dansyl chloride acetone solution (7.5 mg/mL) were added. Next, the mixture was mixed using a vortex and incubated in a heating module for 15 min in the dark at 60 ± 1 °C. Further, 100 μL L-proline (10 mg/mL) was added, shaken manually and the vial was stored in the dark for 15 min. In the next step, 500 μL toluene was added, shaken manually, and stored for 20 min at ≤−18 °C in order to freeze the water phase. Then, as much as possible of the upper organic phase was transferred into a new glass vial and evaporated to dryness under nitrogen at 44 °C. The residue was dissolved in 200 μL (standards in 100 μL) acetonitrile. Before injection into HPLC system the solution was filtered through a 0.2 μm syringe PTFE filter. The sample preparation procedure is shown in Figure 3.
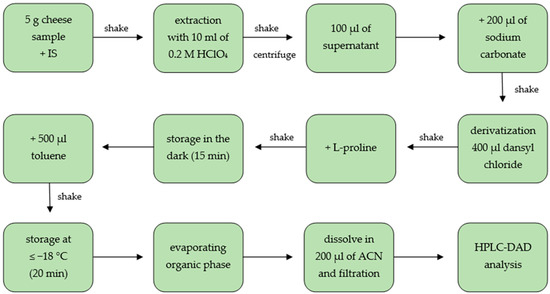
Figure 3. Scheme of sample preparation for the determination of BAs in cheese (IS: internal standard, ACN: acetonitrile, HPLC-DAD: high performance liquid chromatography with diode array detector).
3.5. Method Validation
The method was validated in terms of its linearity, limit of detection (LOD), limit of quantification (LOQ), precision, and recovery. The calibration curves were obtained by plotting the peak-area ratios of the analyte to the IS versus the concentration. Linearity was evaluated in duplicate at five concentrations of the standard mixtures levels: 5, 10, 20, 100, 200 mg/kg. The LOD and LOQ values for all analytes were calculated based on the data from the appropriate calibration curves. LOD was established as 3.3 times the value of residual standard deviation (Sx/y) divided to the slope of the calibration curve (a). LOQ was established as 10 times the (Sx,y) divided to (a). The cheeses samples free from biogenic amines were used as blank, to spike aliquots for validation studied. To assess the accuracy of the developed method, precision and recovery were evaluated. Spiked of the same cheese sample in six replicates for each level analyzed within one working day were used to estimate the intra-day precision. The inter-day precision was assessed for each level based eighteen replicates spiked of the various cheese samples within three working days. The method’s precision was determined by calculating the relative standard deviation (RSDr). Average recovery percentages were calculated by comparison the determined concentrations of spiked samples to their target level.
3.6. Real Samples
The developed and validated method was applied to analyze biogenic amines in 35 samples of ripened cheeses. The investigation covered mold-ripened soft cheeses (Brie, Camembert types), blue-veined cheeses (Roquefort, Gorgonzola types) and hard cheeses (Grana Padano, Parmesan types). Cheeses were purchased from various local retail markets in the Pulawy region (Poland) and the samples were immediately delivered to the laboratory at refrigerated temperature.
“
